AiChemy: Next-generation agent with MCP, skills and custom data for drug discovery
AiChemy accelerates drug discovery by integrating custom and external data (OpenTargets, PubChem, PubMed) through a multi-agent system utilizing MCP, Skills, AI Search, and Genie on Databricks
by Yen Low and Sean Zhang
- A guide to building AiChemy, a multi-agent system on Databricks that integrates external knowledge bases (OpenTargets, PubChem, PubMed) via the Model Context Protocol (MCP) with structured and unstructured data on Databricks.
- The challenge it solves: It accelerates cross-disciplinary drug discovery research by enabling autonomous collaboration between diverse AI agents, allowing them to sift through massive, disparate datasets and provide traceable, evidence-backed findings.
- Outcomes: Researchers can identify disease targets, evaluate drug candidates, retrieve detailed properties, and perform safety assessments, leading to more efficient drug discovery and lead generation.
Multi-agent systems accelerate cross-disciplinary research
Imagine multi-agent AI systems collaborating like a team of cross-disciplinary experts, autonomously sifting through massive datasets to uncover novel patterns and hypotheses. This is now conveniently achievable with Model Context Protocol (MCP), a new standard for easily integrating diverse data sources and tools. The growing MCP server ecosystem—from knowledge bases to report generators—offers endless capabilities.
What AiChemy does
Meet AiChemy, a multi-agent assistant that combines external MCP servers like OpenTargets, PubChem, and PubMed with your own chemical libraries on Databricks such that the combined knowledge bases can be better analyzed and interpreted together. It also has Skills that can be optionally loaded to provide detailed instructions for producing task-specific reports, consistently formatted for research, regulatory, or business needs.
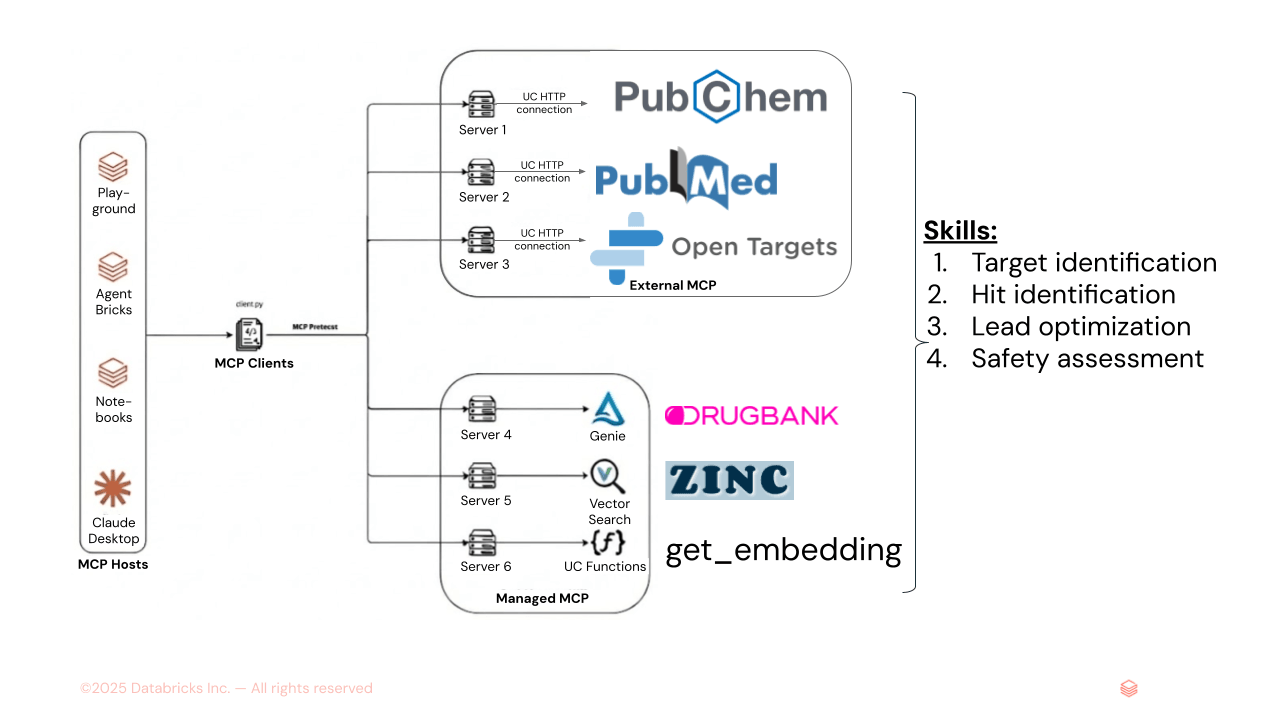
Figure 1. AiChemy is a multi-agent supervisor comprising external MCP servers PubChem, PubMed, and OpenTargets, and Databricks-managed MCP servers of Genie Space (text-to-SQL for DrugBank structured data) and of AI Search (for unstructured data like ZINC molecular embeddings). Skills can also be loaded to specify task sequence and report formatting and style to ensure consistent output.
Its key capabilities include identifying disease targets and drug candidates, retrieving their detailed chemical, pharmacokinetics properties, and providing safety and toxicity assessments. Crucially, AiChemy backs its findings with supporting evidence traceable to verifiable data sources, making it ideal for research.
Use Case 1: Understand disease mechanisms, find druggable targets and lead generation
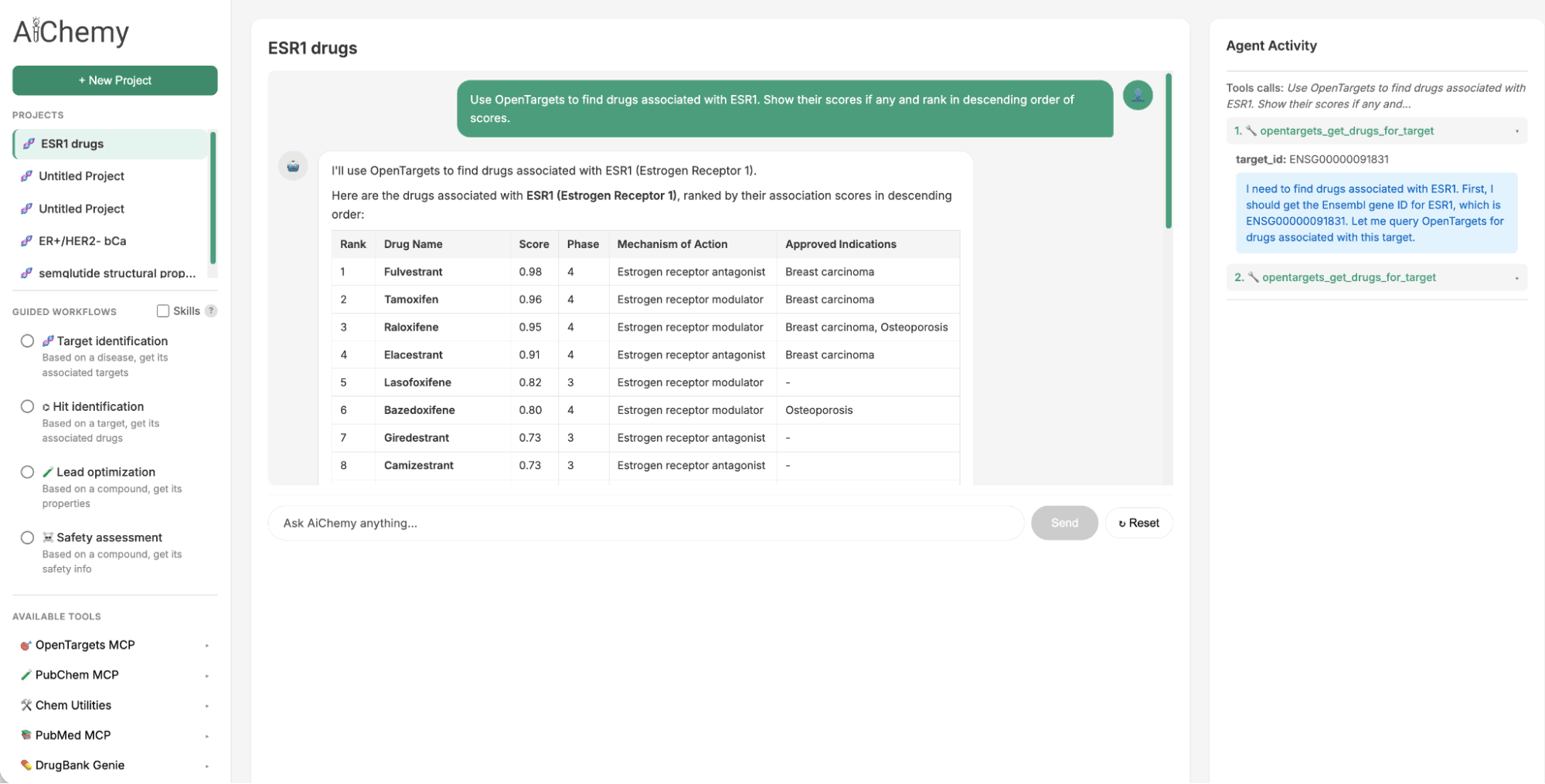
The Guided Tasks panel provides necessary prompts and agent Skills to perform the key steps in a drug discovery workflow of disease -> target -> drug -> literature validation.
- Identify Therapeutic Targets: Starting with a specific disease subtype, such as Estrogen Receptor-positive (ER+)/HER2-negative (HER2-) breast cancer (where ER and HER2 are key protein biomarkers), find associated therapeutic targets (e.g., ESR1).
- Find Associated Drugs: Use the identified target (e.g., ESR1) to find potential drug candidates.
- Validate with Literature: For a given drug candidate (e.g., camizestrant), check the scientific literature for supporting evidence.
Use Case 2: Lead generation by chemical similarity
To identify a follow-up to the oral Selective Estrogen Receptor Degrader (SERD) approved in 2023, Elacestrant, we can leverage chemical similarity. We search the large ZINC15 chemical library for drug-like molecules structurally similar to Elacestrant, as Quantitative Structure–Activity Relationship (QSAR) principles suggest they will share similar properties. This is achieved by querying Databricks AI Search, which uses the 1024-bit Extended-Connectivity Fingerprint (ECFP) molecular embedding of Elacestrant (as query vector) to find the most similar embeddings within ZINC's 250,000-molecule index.
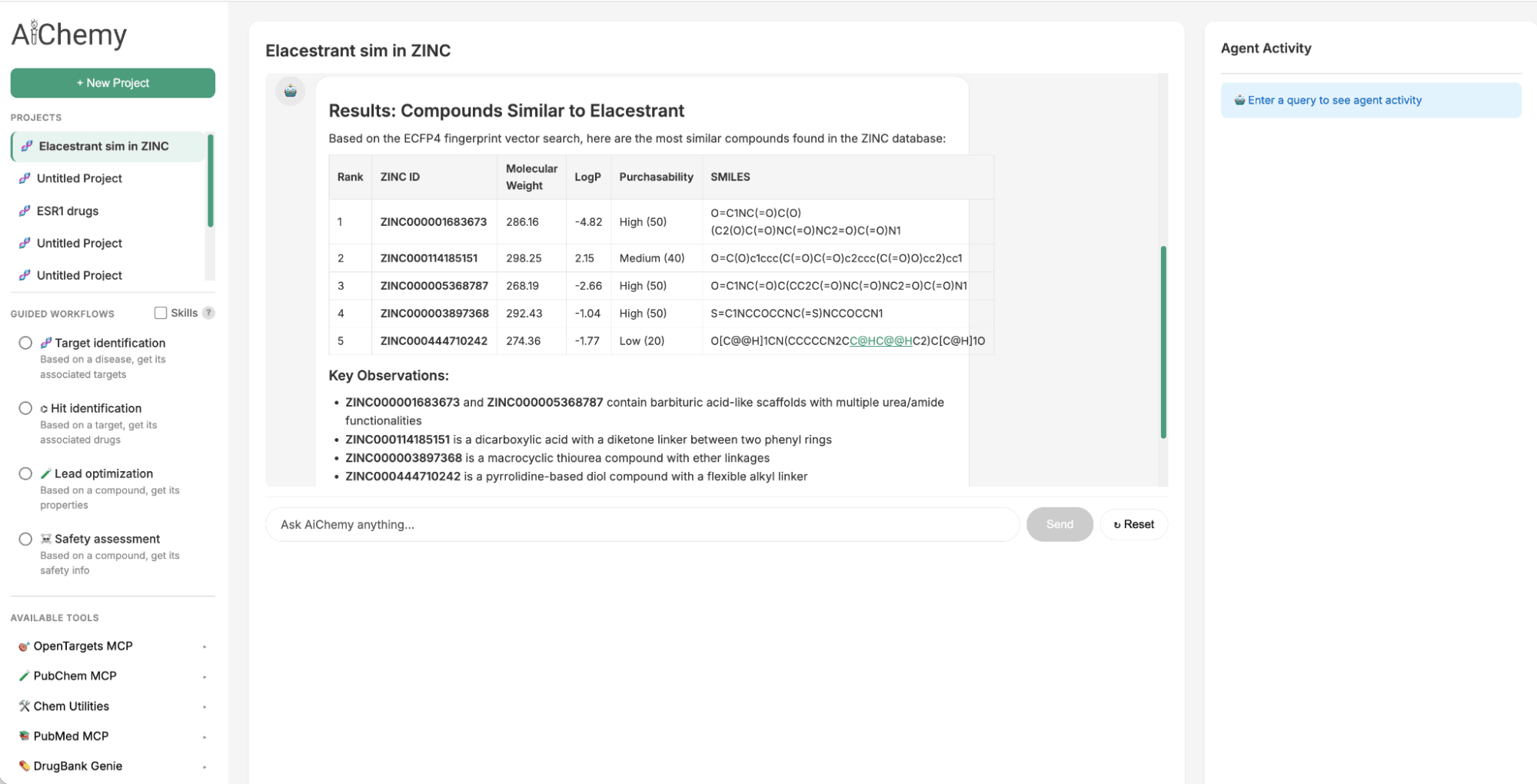
Figure 2. AiChemy includes the vector search of the ZINC database of 250,000 commercially available molecules. This enables us to generate lead compounds by chemical similarity. In this screenshot, we asked AiChemy to find in the ZINC vector search compounds most similar to Elacestrant based on the ECFP4 molecular embedding.
Build your own research multi-agent supervisor
We will customize a multi-agent supervisor on Databricks by integrating public MCP servers with proprietary data on Databricks. To achieve this, you have the option of using either no-code Agent Bricks or coding options like Notebooks. The Databricks Playground allows for quick prototyping and iteration of your agents.
Step 1: Prepare the components required for the multi-agent supervisor
The multi-agent system has 5 workers:
- OpenTargets: external MCP server of a disease-target-drug knowledge graph
- PubMed: external MCP server of biomedical literature
- PubChem: external MCP server of chemical compounds
- Drug Library (Genie): A chemical library with structured drug properties, made into a Genie space to provide text-to-SQL capabilities.
- Chemical Library (AI Search): A proprietary library of unstructured chemical data with molecular fingerprint embeddings, prepared as a vector index to facilitate similarity search by embeddings.
Step 1a: Securely connect to public MCP servers via Unity Catalog (UC) connections in the UI or in a Databricks Notebook (e.g. 4_connect_ext_mcp_opentarget.py).
Step 1b: Ensure your structured table(s) (e.g. DrugBank) is transformed into a Genie space with text-to-SQL functionality using the UI. See 1_load_drugbank and descriptors.py
Step 1c: Ensure your unstructured chemical library is created as a vector index in the UI or in a Notebook to enable similarity search. See 2_create VS zinc15.py
Step 2 (Easy Option): Build the multi-agent supervisor using no-code Supervisor Agent in 2 mins
To assemble them, try the no-code Agent Bricks that builds a supervisor agent with the above components via the UI and deploys it to a REST API endpoint, all in a few minutes.
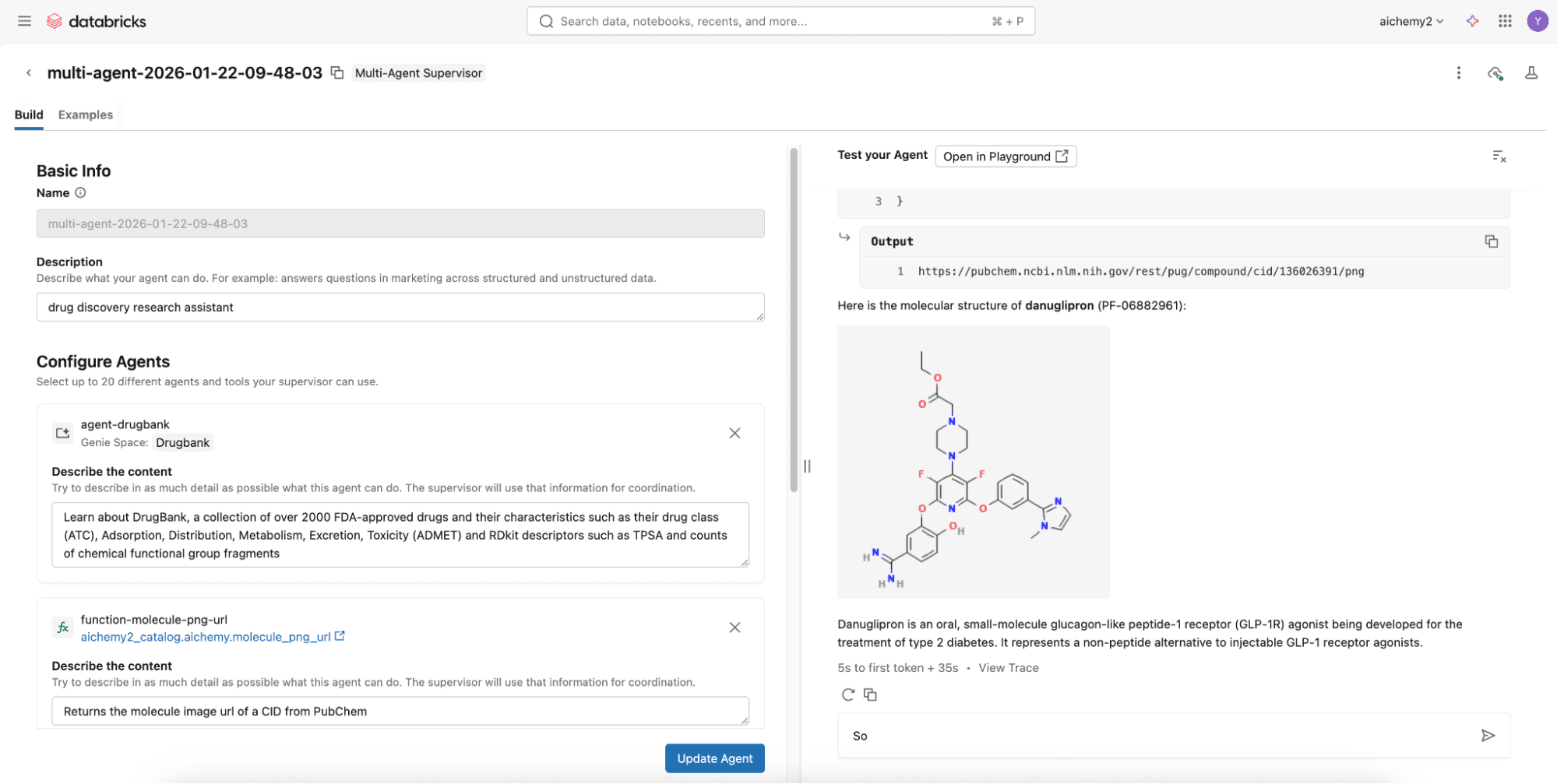
Step 2 (Advanced Option): Build the multi-agent supervisor using Databricks Notebooks
For more advanced capabilities like agentic memory and Skills, develop a Langgraph supervisor on Databricks Notebooks to integrate with Lakebase, Databricks Serverless Postgres database. Check out this code repository where you can simply define the multi-agent components (see Step 1) in the config.yml.
Once config.yml is defined, you can deploy the multi-agent supervisor as a MLflow AgentServer (FastAPI wrapper) with a React web user interface (UI). Deploy them both to Databricks Apps via the UI or Databricks CLI. Set the appropriate permissions for users to use the Databricks App and for the app’s service principal to access the underlying resources (e.g. experiment for logging traces, secret scope if any).
Step 3: Evaluate and monitor your agent
Every invocation to the agent is automatically logged and traced to a Databricks MLflow experiment using OpenTelemetry standards. This enables easy evaluation of the responses offline or online to improve the agent over time. Additionally, your deployed multi-agent uses the LLM behind AI Gateway so you can enjoy the benefits of centralized governance, built-in safeguards, and full observability for production readiness.
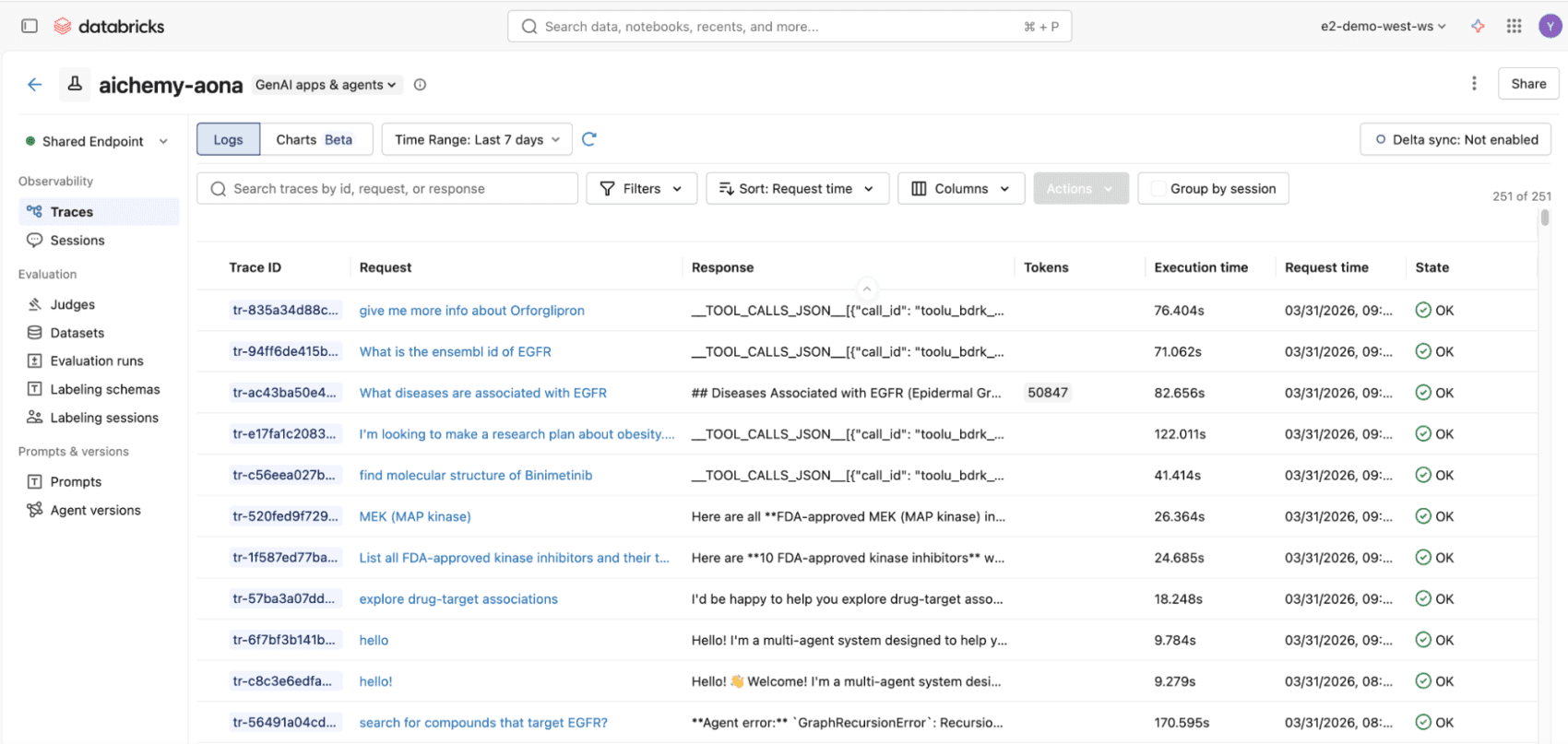
Figure 3. All invocations to the multiagent whether via React UI or REST API will be logged to MLflow traces, compliant with OpenTelemetry standards, for end-to-end observability.
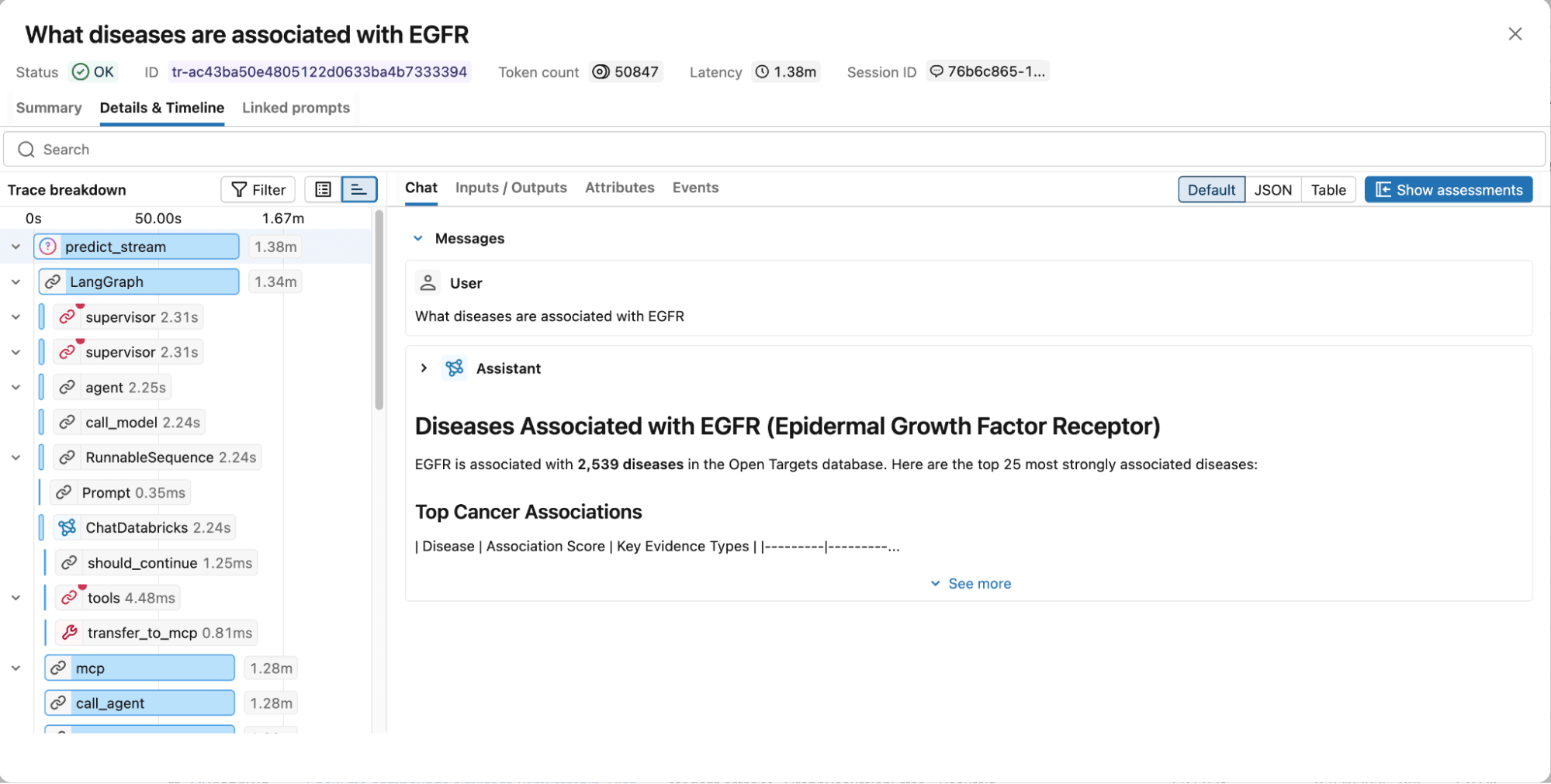
Figure 4. MLflow traces capture the full execution graph, including reasoning steps, tool calls, retrieved documents, latency, and token usage for easy debugging and optimization.
Next Steps
We invite you to explore the AiChemy web app and Github repository. Start building your custom multi-agent system with the intuitive, no-code Agent Bricks framework on Databricks so you can stop sifting and start discovering!

Get the latest posts in your inbox
Subscribe to our blog and get the latest posts delivered to your inbox.